Biomarker Discovery & Validation
Identifying Protein Biomarkers for Diagnosing Paediatric Infectious Disease
Aim
To identify host protein biomarkers for development into an accurate, rapid point-of-care test that can distinguish between bacterial and viral infections in febrile children.
Background & Motivation

Phase 1
Biomarker Discovery
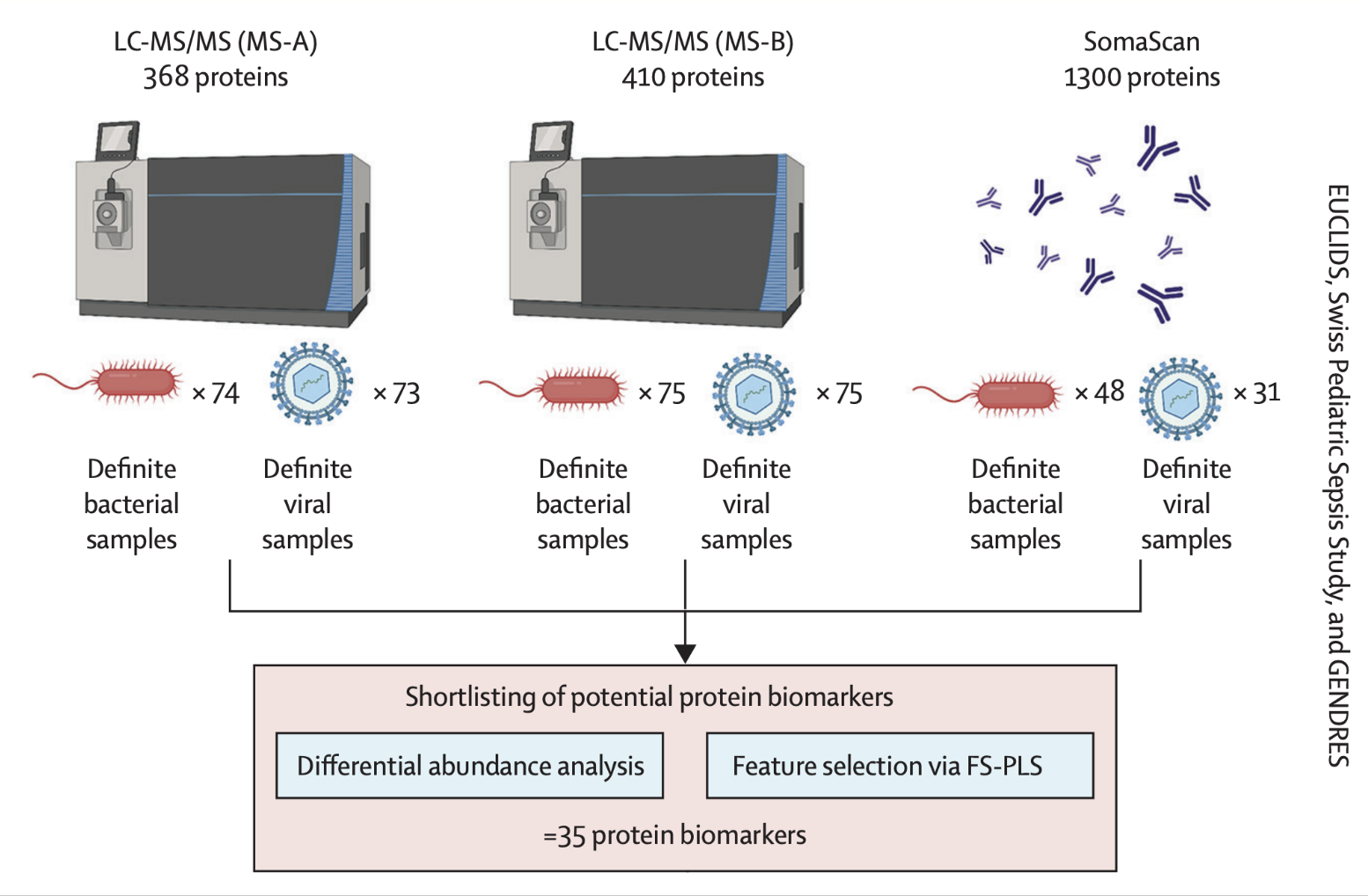
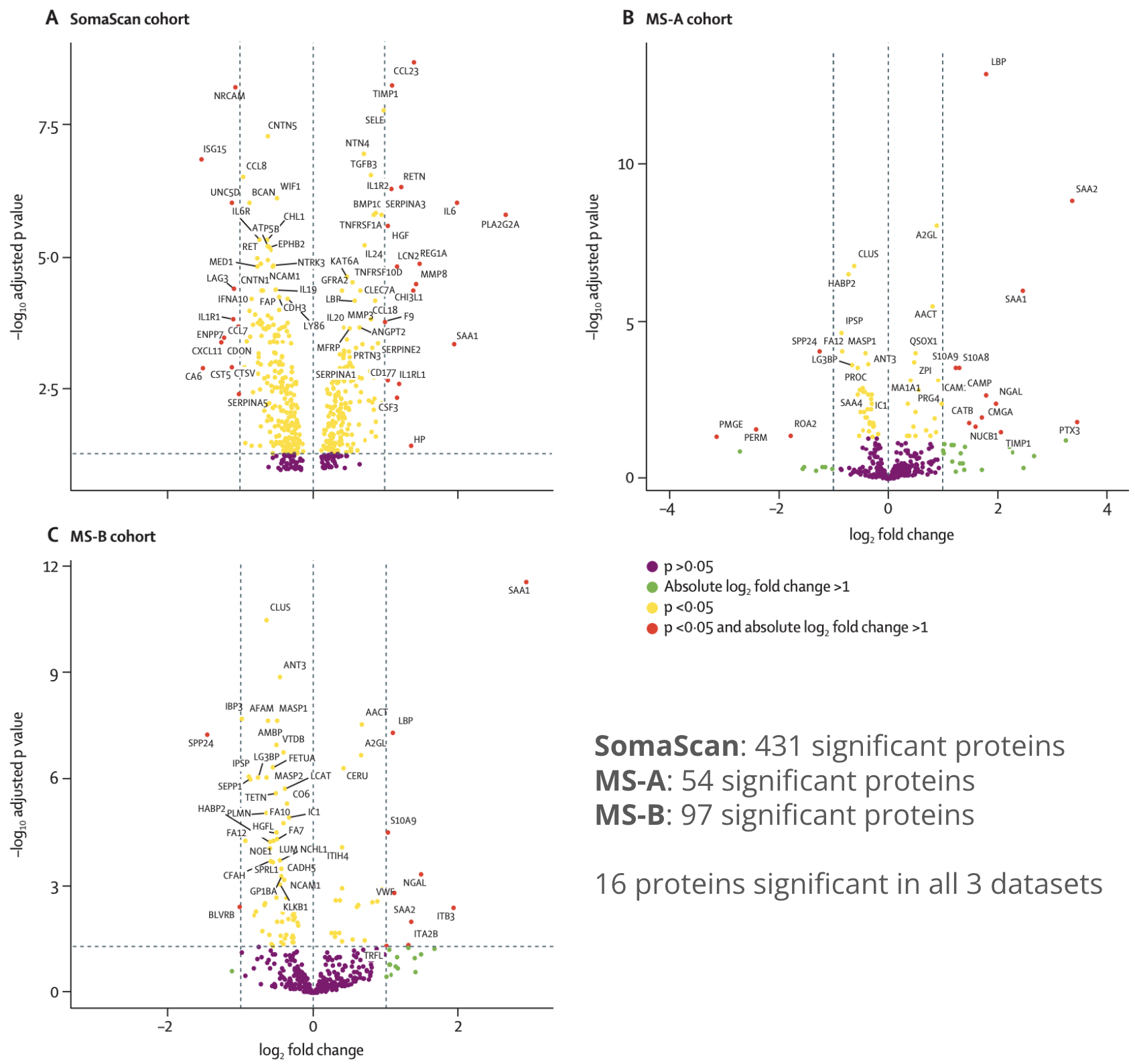
Three high-dimensional proteomic datasets (SomaScan, MS-A, MS-B), each containing confirmed bacterial and viral infection cases, were pre-processed and quality-controlled.
Differential abundance analysis was applied to identify proteins with differing levels between infection types. A machine learning algorithm — forward selection with partial least squares (FS-PLS) combined with repeated cross-validation — was applied to generate hundreds of potential biomarker models. We developed a bespoke ranking method, incorporating both differential abundance analysis and feature selection results to prioritise individual and combinations of biomarkers.
35 protein candidates shortlisted across all 3 datasets
Phase 2
Signature Refinement
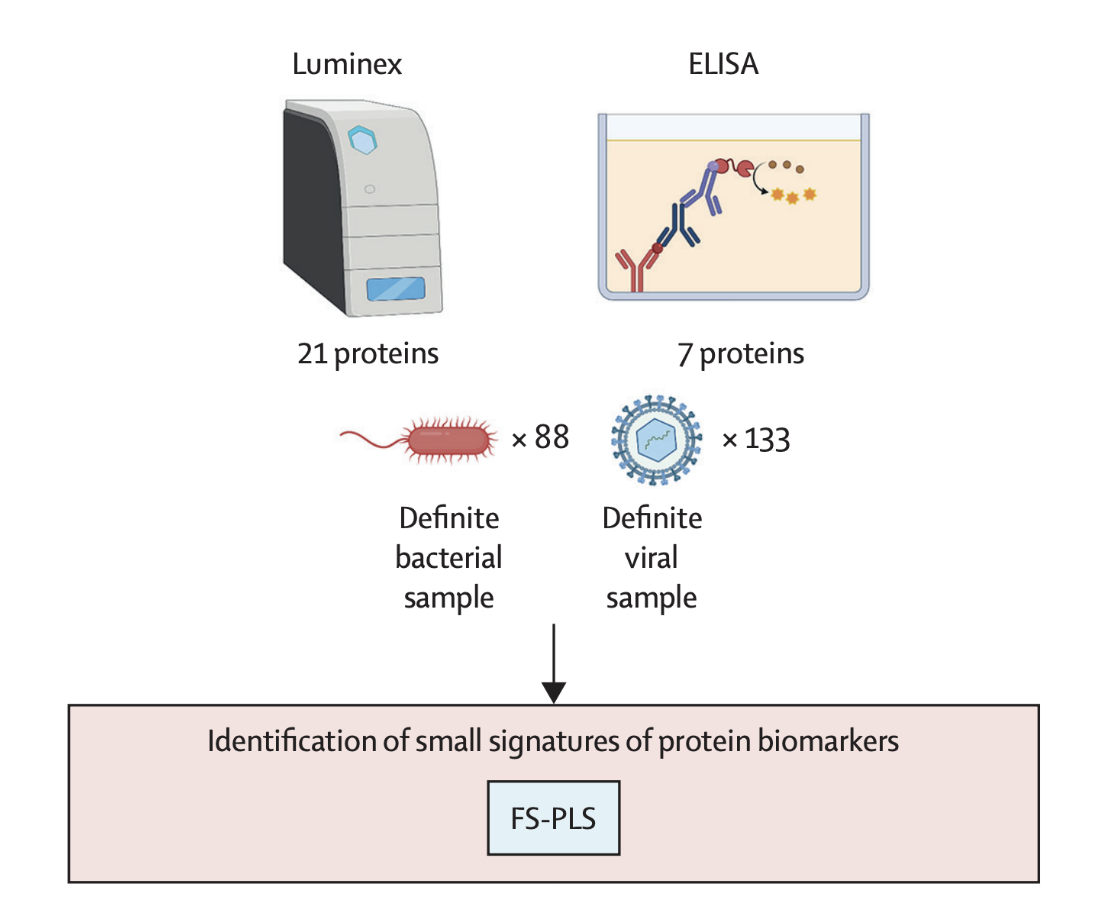
28 of the 35 candidates were measured using ELISA and Luminex immunoassays — low-throughput methods closer to point-of-care methodologies. FS-PLS with repeated cross-validation was reapplied to this refined pool.
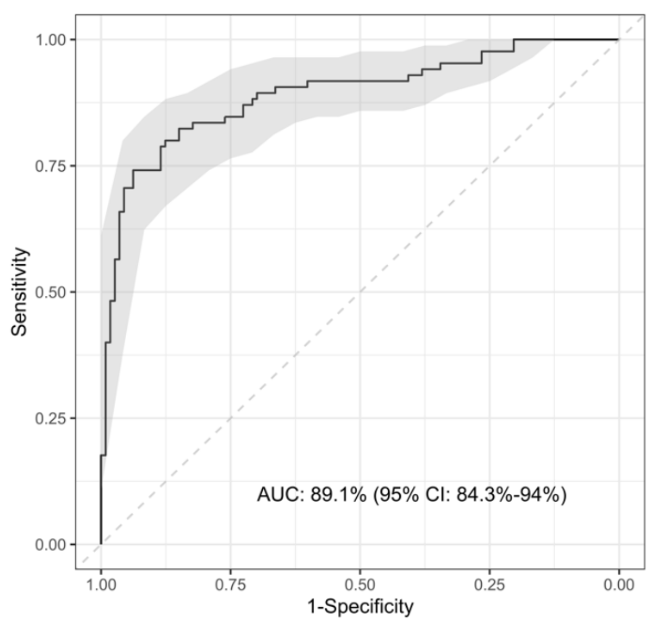
A 5-protein signature was identified — NCAM1, IL-18, SELE, NGAL, and IFN-γ — selected in 7 of 25 iterations with an AUC of 89.1% and with high sensitivity for bacterial infections (90.4%) but lower specificity (67%), mislabelling about one-third of viral cases as bacterial. At this point, we identified Galectin-3BP as an additional marker to improve discrimination between bacterial and viral infections in borderline cases where bacterial infections appeared more viral clinically.
6-protein signature identified
Phase 3
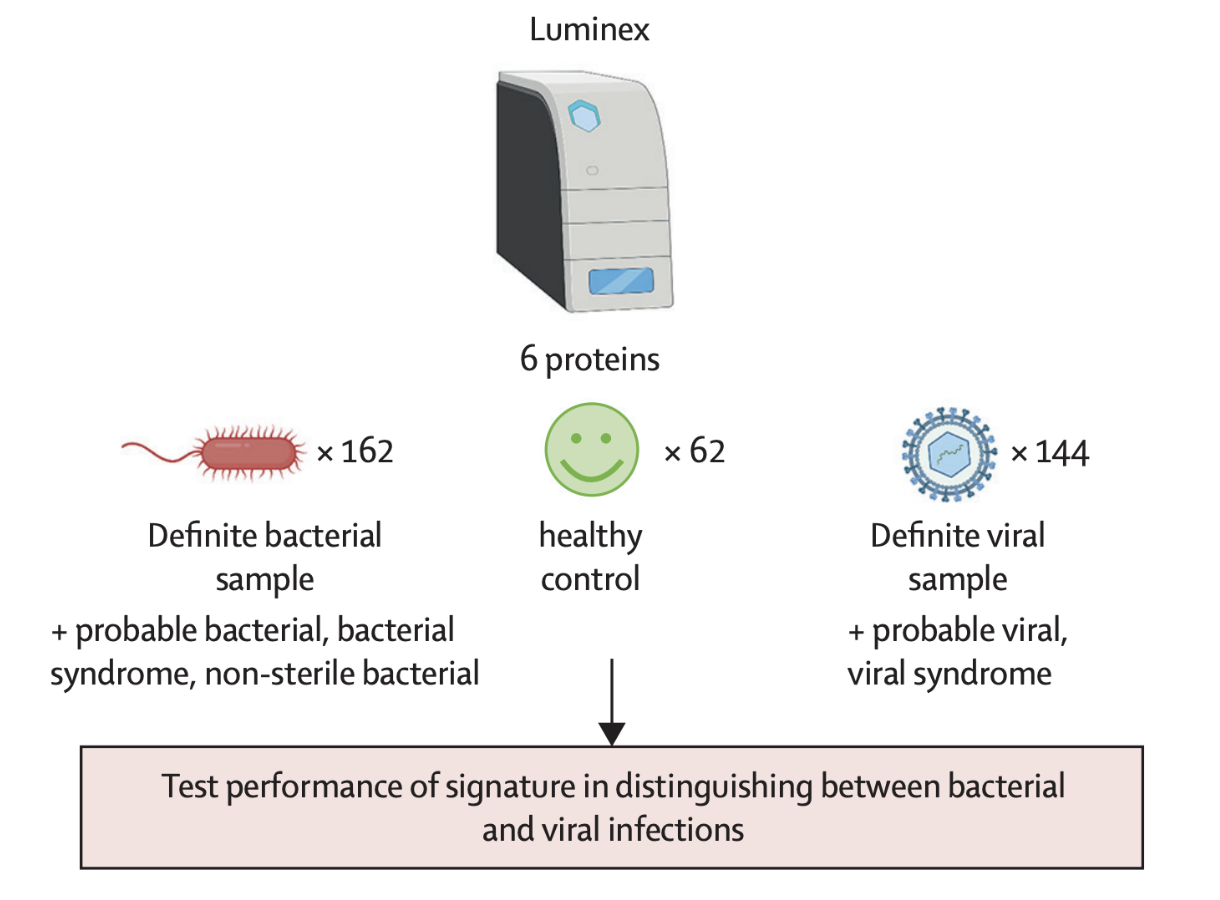
Final Validation
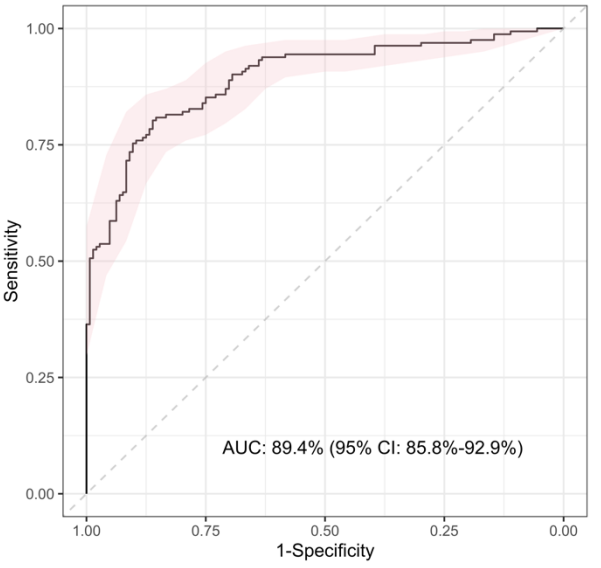
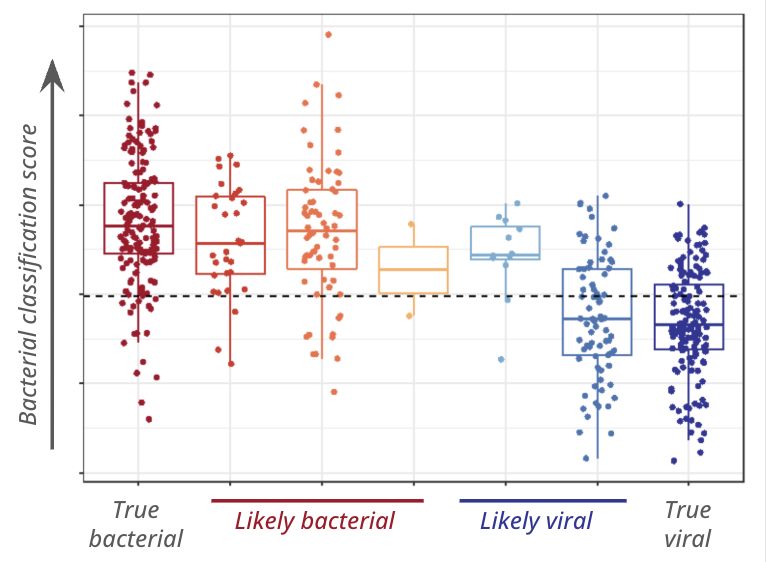
A further validation study was performed using Luminex, this time measuring the 6 optimal proteins identified in phase 2 (NCAM1, IL18, SELE, NGAL, IFN-γ, Galectin-3BP). The 6-protein signature performed well overall (AUC >89%) and could accurately identify both bacterial (sensitivity: 88.1%) and viral (specificity: 84.9%) infections.
Signature discovery used confirmed cases, but most real-world cases are unconfirmed. The signature aligned with clinical assessment even in unconfirmed cases, demonstrating potential to support confident treatment decisions across the full spectrum of childhood fevers.
AUC >89% · Sensitivity 88.1% · Specificity 84.9%
Outcomes
Publication
Published in Lancet Digital Health (Impact Factor 24.1) — a leading peer-reviewed journal in digital medicine.
Patent Filed
A host protein signature for distinguishing between bacterial and viral infections in febrile children. PCT/GB2024/052214, filed August 2024. Jackson HR, Kaforou M, Levin M, Kuijpers T, de Jonge M.
Ongoing Translation
Active work underway at Imperial College London to translate the 6-protein signature into a rapid point-of-care diagnostic test.